This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Proteomics? [1]
|
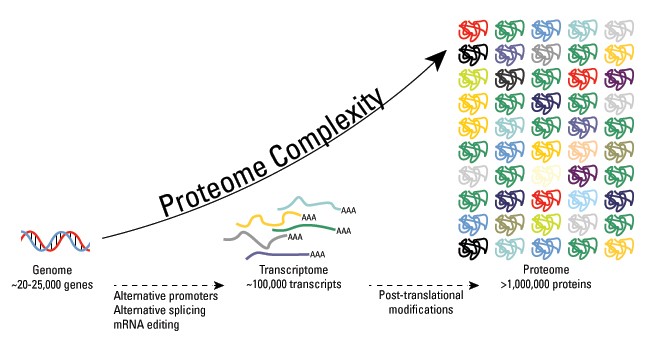
Proteomics is the large-scale study of proteomes, or the sets of proteins produced in an organism or present in a specific sample. Proteomics can be used to determine where proteins are expressed, how they are modified, the movement in the cell, and how proteins interact with each other. By determining this information, proteomics can provide information for many biological problems such as protein interactions in diseased states or if they are involved in a specific process. However, the proteome is complex and dynamic as proteins are constantly changed over time. Post translational modifications attribute to this complexity and explain the challenges when studying proteins.
|
Figure 1: Dynamic Nature and Complexity of the Proteome
|
What are post-translation modifications? [2]
|
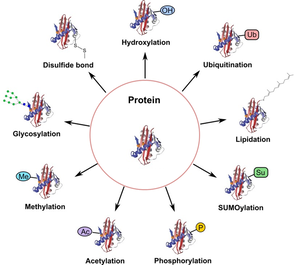
Figure 2: Types of Post-translation Modifications
|
Protein post-translational modifications increase the complexity of the proteome by covalently adding functional groups, proteins, cleaving subunits, or degrading proteins. These modifications include phosphorylation, glycosylation, ubiquitination, methylation, and proteolysis. These modifications lead to normal cell biology. However, their disruption can lead to pathogenesis. Therefore, by understanding the locations and function of these modifications, more knowledge can be obtained on the mechanism of disease and potential prevention.
|
How conserved are FUS phosphorylation sites?
|
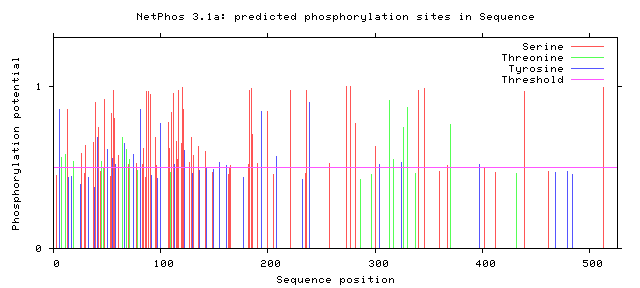
Figure 3: Potential Phosphorylation Sites of Human FUS
|
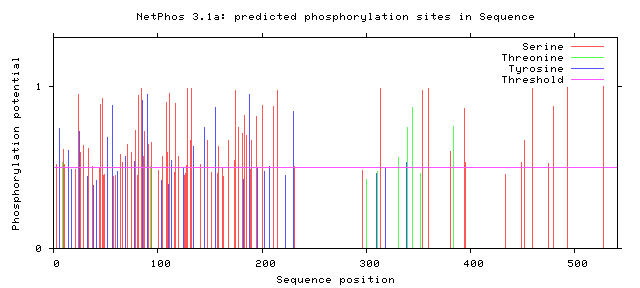
Figure 4: Potential Phosphorylation Sites of Zebrafish FUS
|
Where are phosphorylation sites located in the FUS protein?
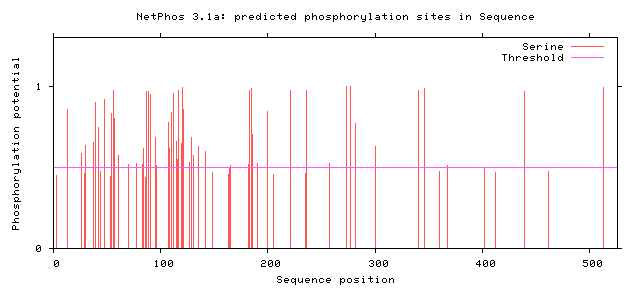
Human FUSFigure 5: Serine Phosphorylation Sites of Human FUS
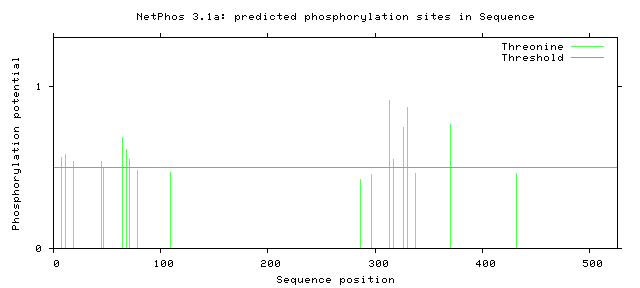
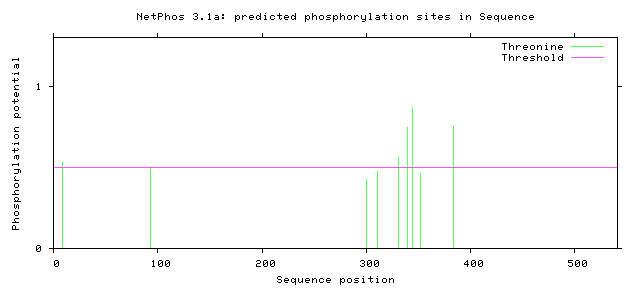
Figure 7: Threonine Phosphorylation Sites of Human FUS
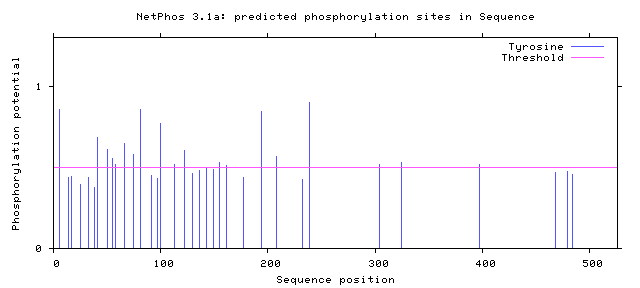
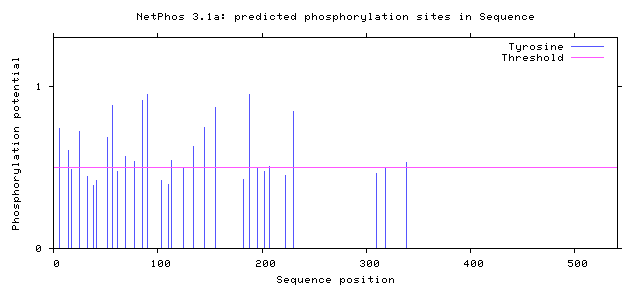
Figure 9: Tyrosine Phosphorylation Sites of Human FUS
|
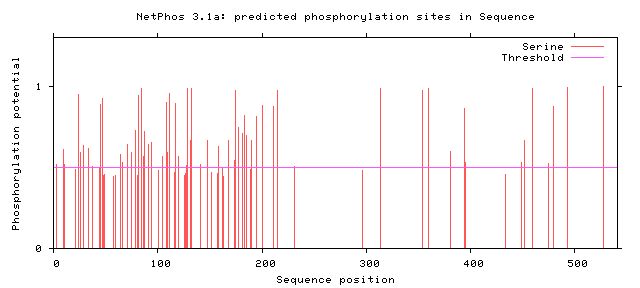
Zebrafish FUSFigure 6: Serine Phosphorylation Sites of Zebrafish FUS
Figure 8: Threonine Phosphorylation Sites of Zebrafish FUS
Figure 10: Tyrosine Phosphorylation Sites of Zebrafish FUS
|
Additional Proteomic Information on FUS [3,4,5]
Isoelectric Point/Molecular Weight: 9.40 / 53425.84
Number of Trypsin Cleavage Sites: 49 potential sites
Transmembrane Protein: No
FUS Subcellular Location: Nucleus
CDC5L Interactor Subcellular Location: Nucleoplasm
CHEK1 Interactor Subcellular Location: Nuceloplasm, Vesicles
BRAC1 Interactor Subcellular Location: Nucleus
Number of Trypsin Cleavage Sites: 49 potential sites
Transmembrane Protein: No
FUS Subcellular Location: Nucleus
CDC5L Interactor Subcellular Location: Nucleoplasm
CHEK1 Interactor Subcellular Location: Nuceloplasm, Vesicles
BRAC1 Interactor Subcellular Location: Nucleus
Conclusion
From these findings, there were two potential phosphorylation sites in the zinc finger domain and fourteen potential sites present in the RNA recognition motif. Only two human variants matched phosphorylation sites on the C terminal, but almost all phosphorylation sites near the N terminal matched known human variant. The residue at amino acid 526 has already been shown to impact nuclear transport [6]. The phosphorylation sites at the low complexity regions near the N terminal have also already been shown to facilitate aggregation [7]. However, no work has looked at altering the phosphorylation sites suggested in the Zinc finger domain and RNA recognition motif. In addition, researchers have noticed phosphorylation of FUS in response to DNA damage in the N terminus. However, researchers have failed to look at alterations to these sites and the effect on DNA repair. These potential phosphorylation sites could be interesting to alter and to discover which sites are critical for DNA repair.
References
1. Proteomics: An introduction to EMBL-EBI resources. (2019). Retrieved on 4/11/2019 from https://www.ebi.ac.uk/training/online/course/proteomics-introduction-ebi-resources/what-proteomics
2. Overview of Post-Translational Modifications. (2019). Retrieved on 4/11/2019 from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
3. Retrieved on 4/11/2019 from https://web.expasy.org/peptide_cutter/
4. Retrieved on 4/11/2019 from http://expasy.org/tools/pi_tool.html
5. Retrieved on 4/11/2019 from http://www.cbs.dtu.dk/services/TMHMM-2.0/
6. Monahan, Z., Ryan, V. H., Janke, A. M., Burke, K. A., Rhoads, S. N., Zerze, G. H., … Fawzi, N. L. (2017). Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. The EMBO journal, 36(20), 2951–2967. doi:10.15252/embj.201696394
7. Darovic, S., Mihevc, S.P., Zupunski, V. et al. (2015, September). Phosphorylation of C-terminal tyrosine residue 526 in FUS impairs its nuclear import. Journal of Cell Science, 128: 4151-4159. doi: 10.1242/jcs.176602
Header Image: https://images.fineartamerica.com/images-medium-large-5/13-animal-cell-organelles-artwork-science-photo-library.jpg
Figure 1: https://www.proteomicscenter.nl/background-information/
Figure 2: https://www.creative-proteomics.com/blog/wp-content/uploads/2018/06/Strategies-for-Post-Translational-Modifications-PTMs-0.png
Figure 3-10: Hyperlinked to http://www.cbs.dtu.dk/services/NetPhos/
2. Overview of Post-Translational Modifications. (2019). Retrieved on 4/11/2019 from https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
3. Retrieved on 4/11/2019 from https://web.expasy.org/peptide_cutter/
4. Retrieved on 4/11/2019 from http://expasy.org/tools/pi_tool.html
5. Retrieved on 4/11/2019 from http://www.cbs.dtu.dk/services/TMHMM-2.0/
6. Monahan, Z., Ryan, V. H., Janke, A. M., Burke, K. A., Rhoads, S. N., Zerze, G. H., … Fawzi, N. L. (2017). Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. The EMBO journal, 36(20), 2951–2967. doi:10.15252/embj.201696394
7. Darovic, S., Mihevc, S.P., Zupunski, V. et al. (2015, September). Phosphorylation of C-terminal tyrosine residue 526 in FUS impairs its nuclear import. Journal of Cell Science, 128: 4151-4159. doi: 10.1242/jcs.176602
Header Image: https://images.fineartamerica.com/images-medium-large-5/13-animal-cell-organelles-artwork-science-photo-library.jpg
Figure 1: https://www.proteomicscenter.nl/background-information/
Figure 2: https://www.creative-proteomics.com/blog/wp-content/uploads/2018/06/Strategies-for-Post-Translational-Modifications-PTMs-0.png
Figure 3-10: Hyperlinked to http://www.cbs.dtu.dk/services/NetPhos/