This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are Protein Domains? [1]
Protein domains are independent genetic segments that over evolution have formed a single-domain protein or multi-domain protein. The domain can have an independent purpose or can coordinate with another domain to help function as a multi-domain protein. Proteins can be formed by duplication, divergence and recombination of domains. These domains are important in determining the processes with the protein and help suggest mechanisms associated with a certain disease.
Protein Domains of FUS [2]
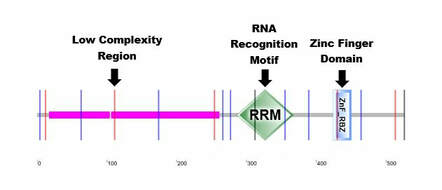
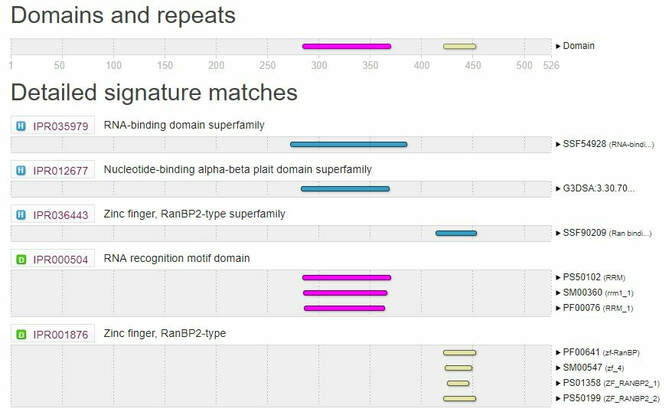
Utilizing search tools of SMART, Pfam, InterPRo, and Prosite, it was revealed that FUS contained two important protein domains. The first of which, is a 78 amino acid RNA Recognition Motif abbreviated as RRM1 that is important for ssRNA binding. The second is a 131 amino acid Zinc finger domain abbreviated as ZfRanBP, which is important for nuclear transport and other cellular processes. Due to consistency across all sites, it was suggested that these results are accurate and for accurate comparison between other species.
Figure 3: InterPro Domains of FUS
What Biological Role Does Each Protein Domain Play?
RNA Recognition Motif [3]
|
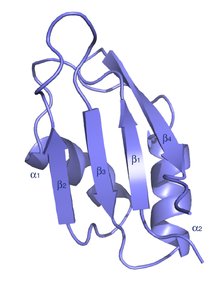
Figure 4: Schematic Depiction of the RNA Binding Motif
|
An RNA recognition motif is a RNA-binding domain approximately 90 amino acids in length that bind ssRNAs and is present in many different types of eukaryotic protein structures. The largest group of these binding domains are the RNA recognition motif (RRM) family and contain an eight amino acid RNP-1 sequence. These RRM proteins are implicated in regulating alternative splicing, RNA stability, and translation. RRM proteins usually contain four anti-parallel beta-strands and two alpha-helices that arrange themselves in a beta and alpha alternating pattern. This alternating patterns helps to create a folded protein with side chains.
|
Zinc Finger Domain [4]
|
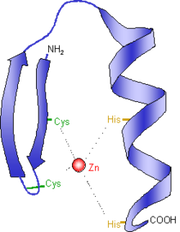
Zinc finger (Znf) domains are stable small protein motifs that contain finger-like projections to bind with their target and rarely undergo conformation changes. Many of these domains bind with zinc, but some can bind to other metals and nonmetals. These domains are important for DNA, RNA, protein, and lipid binding. These binding relationships depend on the specific sequence, links between finger projections, higher-order structures, and the specific amount of projections. Proteins containing a Znf region function in gene transcription, translation, mRNA trafficking, cytoskeleton organization, and many other cellular processes.
The zinc finger domain in FUS is similar to those found in RanBP2 proteins. Ran is a conserved member of Ras superfamily that is involved with receptor-mediated transport between the nucleus and cytoplasm. Showing RanBP2 plays a critical role in nuclear protein import. |
Figure 5: Schematic Diagram of the Zinc Finger Domain
|
Which Domains of FUS Are Conserved?
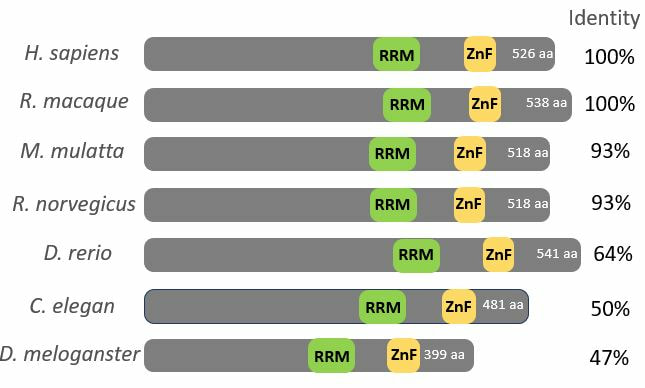
Upon analysis of FUS via Smart, Pfam, InterPRo, and Prosite, it was shown that the RNA Recognition Motif, and Zinc Finger Domain is conserved among model organisms shown in Figure 3. These two domains are critical for the function of the protein and explains why there are very few differences of these domains between species. In addition to these domains, there also low complexity regions in the C terminal region of the protein across all species. This low complexity region is simply repeated amino acids and may be involved with various process. Recent research has shown that this region participates in liquid-to-phase transition in humans [5].
Figure 6: Conserved RNA binding and Zinc Finger Domains Across Species
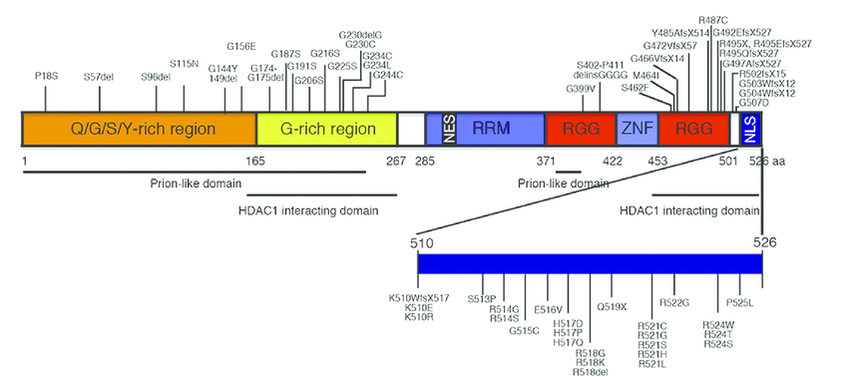
Where Are FUS Mutations Located? [6]
Figure 7: Regions Where FUS Mutations Occur
Conclusion
The results from protein domains suggested by these program showed that these two domains were well conserved across several species. These two domains were RNA recognition motif domain and Zinc finger domain. Each domain was conserved across homologs implying that each domain plays a critical role in binding and the function of the protein. It is also interesting to note that no mutations strictly associated with ALS develop in these two domains. This suggests that both of these domains are critical for RNA binding and other roles that this protein might participate in.
References
1. Vogel, C., Bashton, N., et. al. (2004, April). Structure, function and evolution of multidomain proteins. Current Opinion in Structural Biology,14:208–216 DOI:10.1016/j.sbi.2004.03.011
2. Protein: FUS_HUMAN (P35337). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/protein/P35637.
3. Family: RRM_1 (PF00076). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/family/PF00076.
4. Family: zf-RanBP (PF00641). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/family/PF00641.
5. Patel, A., Lee, H.O., et al. (2015, August 27). A Liquid-to-Solid Phase Transition of the ALS Protein FUS Accelerated by Disease Mutation. Cell, 162(5): 1066-1077. DOI:https://doi.org/10.1016/j.cell.2015.07.047
6. Shang, Y., Huang, E.J. (2016, September). Mechanisms of FUS mutations in familial amyotrophic lateral sclerosis
Brain Research, 1647:65-78. DOI: 10.1016/j.brainres.2016.03.036
Figure 1: Original Image from http://smart.embl-heidelberg.de/smart/show_motifs.pl
Figure 2: Original Image from http://pfam.xfam.org/protein/P35637
Figure 3: https://www.ebi.ac.uk/interpro/sequencesearch/iprscan5-S20190303-012251-0973-40831918-p1m
Figure 4: http://pfam.xfam.org/family/PF00076
Figure 5: https://upload.wikimedia.org/wikipedia/commons/thumb/f/f2/Zinc_finger.png/220px-Zinc_finger.png
Figure 6: Created by Nathan Johnson
Figure 7: https://www.sciencedirect.com/science/article/pii/S0006899316301652?via%3Dihub
2. Protein: FUS_HUMAN (P35337). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/protein/P35637.
3. Family: RRM_1 (PF00076). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/family/PF00076.
4. Family: zf-RanBP (PF00641). (2019, March). Retrieved March 3, 2019 from http://pfam.xfam.org/family/PF00641.
5. Patel, A., Lee, H.O., et al. (2015, August 27). A Liquid-to-Solid Phase Transition of the ALS Protein FUS Accelerated by Disease Mutation. Cell, 162(5): 1066-1077. DOI:https://doi.org/10.1016/j.cell.2015.07.047
6. Shang, Y., Huang, E.J. (2016, September). Mechanisms of FUS mutations in familial amyotrophic lateral sclerosis
Brain Research, 1647:65-78. DOI: 10.1016/j.brainres.2016.03.036
Figure 1: Original Image from http://smart.embl-heidelberg.de/smart/show_motifs.pl
Figure 2: Original Image from http://pfam.xfam.org/protein/P35637
Figure 3: https://www.ebi.ac.uk/interpro/sequencesearch/iprscan5-S20190303-012251-0973-40831918-p1m
Figure 4: http://pfam.xfam.org/family/PF00076
Figure 5: https://upload.wikimedia.org/wikipedia/commons/thumb/f/f2/Zinc_finger.png/220px-Zinc_finger.png
Figure 6: Created by Nathan Johnson
Figure 7: https://www.sciencedirect.com/science/article/pii/S0006899316301652?via%3Dihub