This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein interaction networks? [1]
Protein-protein interaction networks are mathematical representations of the contacts between proteins present in a cell or sample. These connections are specific, occur along biding regions, and have a specific biological meaning. These interaction networks can highlight stable interactions present in protein complexes and transient interactions like protein kinases or nuclear pore imports. By understanding these protein interaction networks, unknown proteins can be assigned roles, steps in a signaling pathway can be detailed, and the relationship between proteins can be identified.
The interactome is also the summation of all protein networks in a cell or specific sample. Mass-spectrometry, yeast two-hybrid screens, and co-precipitation can provide evidence of protein-protein interactions. With all this complex data, algorithms are needed to sort through potential background and highlight true protein-protein interactions. Various databases like IntAct, Biogrid, and String can provide information on experimental molecular interactions present in the database.
The interactome is also the summation of all protein networks in a cell or specific sample. Mass-spectrometry, yeast two-hybrid screens, and co-precipitation can provide evidence of protein-protein interactions. With all this complex data, algorithms are needed to sort through potential background and highlight true protein-protein interactions. Various databases like IntAct, Biogrid, and String can provide information on experimental molecular interactions present in the database.
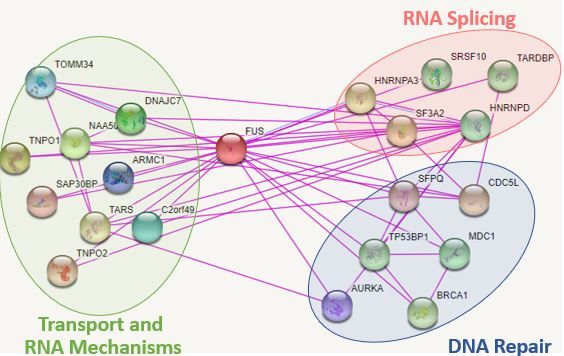
FUS Interactions in Humans by GO Terms:
Figure 3: Interaction network of FUS in humans
Additional interactions not shown: CHEK1 [3]
Conclusion
From these interaction databases, I found that many of the protein interactions of FUS have been associated with cellular transport, RNA metabolism, and RNA splicing as expected. I also found that FUS has interactions with various genes involved with genomic stability like cell cycle checkpoint inhibitors. More interestingly, I found that FUS interacts with CDC5L in both humans and zebrafish [2]. This gene is believed to also play a role in the response to DNA damage leading to DNA repair. However, very little research has looked into this interaction and role in DNA damage. STRING and IntAct provided interactions with BRCA1 and CHEK1 associated with DNA repair, which have been previously mentioned to localize to transcription related DNA repair [3,4]. By understanding the understanding protein-protein interactions of FUS, researchers can highlight the true mechansim of FUS in DNA damage and DNA repair.
References
1. Protein-protein interaction networks. (2019). Retrieved on 4/11/2019 from https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
2. SMART. Retrieved on 4/11/2019 from https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
3. IntAct. Retrieved on 4/11/2019 from https://www.ebi.ac.uk/intact/home.xhtml?status=exp
4. Hill, S.J., Mordes, D.A., Cameron, L.A. et. al. (2016, November). ALS genes in transcription-associated DNA damage.
Proceedings of the National Academy of Sciences. 113(48) DOI:10.1073/pnas.1611673113
Header Image: https://cdn-images-1.medium.com/max/1600/1*xJ7ew1IqICMwkqKNwD0J-w.jpeg
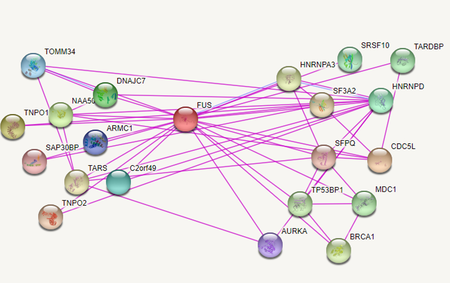
Figure 1: https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
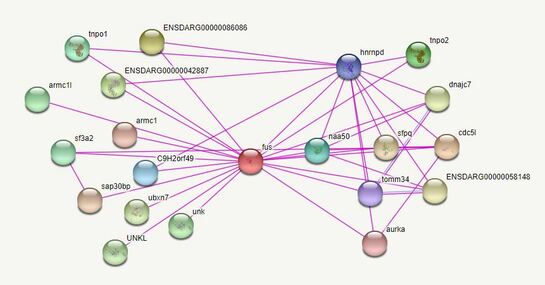
Figure 2: https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
Figure 3: https://string-db.org/cgi/network.pl?taskId=md44EaGXe13x
2. SMART. Retrieved on 4/11/2019 from https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
3. IntAct. Retrieved on 4/11/2019 from https://www.ebi.ac.uk/intact/home.xhtml?status=exp
4. Hill, S.J., Mordes, D.A., Cameron, L.A. et. al. (2016, November). ALS genes in transcription-associated DNA damage.
Proceedings of the National Academy of Sciences. 113(48) DOI:10.1073/pnas.1611673113
Header Image: https://cdn-images-1.medium.com/max/1600/1*xJ7ew1IqICMwkqKNwD0J-w.jpeg
Figure 1: https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
Figure 2: https://version11.string-db.org/cgi/input.pl?sessionId=Odz7uTVCbz7d&input_page_show_search=on
Figure 3: https://string-db.org/cgi/network.pl?taskId=md44EaGXe13x