This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Homology?
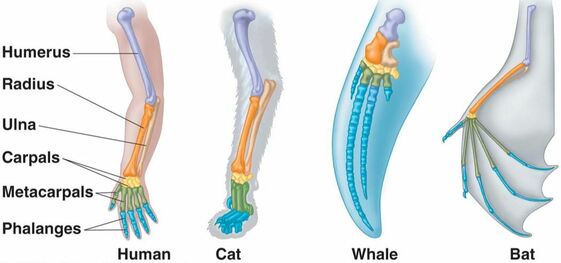
Homology is defined as the relationship between two organisms that are linked to a common descent through some form of identity [1]. These can be relations through physical characteristics, cellular mechanisms, or genetic sequences. An example of homology can be shown in the figure below showing that many species have homologous bone structures and show evidence of a common ancestor (Figure 1). In addition, genetic sequences can be used to determine homology and the similar genetic regions between two different organisms can be referred to as homologs.
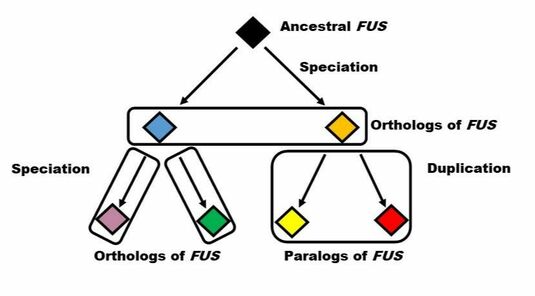
Homologs encompass two sub types that can be broken down into orthologs and paralogs. Orthologs are similar genes in two different species that were derived from a common ancestor. Paralogs are corresponding genes in one species that were derived from a gene duplication during the ancestral lineage of the organism. The figure below shows these two divisions and how an ancestral copy of FUS lead to both orthologs and paralogs of FUS (Figure 2).
Protein homology is also one way of comparing genetic similarities between species. Protein homology can be determined by utilizing programs like BLAST and FASTA to provide protein structures and relatedness among species on the genetic level. Below you can find my homologs for the FUS protein. Each homolog provides the protein name in that species, size of the protein, accession number, and percentage of similarity between humans.
Homologs encompass two sub types that can be broken down into orthologs and paralogs. Orthologs are similar genes in two different species that were derived from a common ancestor. Paralogs are corresponding genes in one species that were derived from a gene duplication during the ancestral lineage of the organism. The figure below shows these two divisions and how an ancestral copy of FUS lead to both orthologs and paralogs of FUS (Figure 2).
Protein homology is also one way of comparing genetic similarities between species. Protein homology can be determined by utilizing programs like BLAST and FASTA to provide protein structures and relatedness among species on the genetic level. Below you can find my homologs for the FUS protein. Each homolog provides the protein name in that species, size of the protein, accession number, and percentage of similarity between humans.
Figure 1: Homologous Structures
Figure 2: Ortholog and Paralog Example in FUS
FUS Protein Homologs
Homo sapiens
|
Pan troglodytes
|
Macaca mulatta
|
| FUS Text File | |
| File Size: | 6 kb |
| File Type: | txt |
Conclusion
The FUS protein found in humans has been shown to have homologs in many different organisms with nervous systems. Knowing the important regulatory roles FUS plays in larger complex organisms, it is no surprise that one would see higher conserved regions for those organisms. Knowing this homology, it is important to also look at the phylogenetics to determine how these homologs have evolved over time. Further studies of the relationship between sequence and function of FUS across species would be important for accessing the major role underlying ALS.
To analyze this specific relationship between sequence and function, the project will use D. rerio (zebrafish) as a model organism. The zebrafish contains a homolog with a relatively conserved protein sequence compared to humans. These zebrafish contain a similar CNS and PNS nervous system and early developmental gene expression that would be important for analyzing motor neuron structure and contain applicable aspects to humans [2]. This could serve for analyzing specific amino acid substitutions found in humans and looking for those associated with that phenotype.
To analyze this specific relationship between sequence and function, the project will use D. rerio (zebrafish) as a model organism. The zebrafish contains a homolog with a relatively conserved protein sequence compared to humans. These zebrafish contain a similar CNS and PNS nervous system and early developmental gene expression that would be important for analyzing motor neuron structure and contain applicable aspects to humans [2]. This could serve for analyzing specific amino acid substitutions found in humans and looking for those associated with that phenotype.
References
1. Koonin, E.V. (2005, August). Orthologs, Paralogs, and Evolutionary Genomics. Annual Review of Genetics, 39:309–338. DOI:10.1146/annurev.genet.39.073003.114725
2. Guo, S. (2009, July). Using zebrafish to assess the impact of drugs on neural development and function. Expert Opinion Drug Discovery, 4(7): 715–726. DOI:10.1517/17460440902988464
Figure 1: http://2.bp.blogspot.com/-AbPRwZgBqFM/USA1Il0yeRI/AAAAAAAAAAk/-NqXfRoaN54/s1600/homologous_forelimbs.jpg
Figure 2: Created by Nathan Johnson
2. Guo, S. (2009, July). Using zebrafish to assess the impact of drugs on neural development and function. Expert Opinion Drug Discovery, 4(7): 715–726. DOI:10.1517/17460440902988464
Figure 1: http://2.bp.blogspot.com/-AbPRwZgBqFM/USA1Il0yeRI/AAAAAAAAAAk/-NqXfRoaN54/s1600/homologous_forelimbs.jpg
Figure 2: Created by Nathan Johnson