This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What Are Model Organisms? [1]
Model organisms are life forms that serve as tools for understanding the biology of human diseases and processes. These species can be easily manipulated and have sequenced genomes in order to analyze specific regions of interest. In addition, these tend to be easy to maintain and can have numerous experimental advantages. There are multiple organisms that have been utilized as model organisms including yeast, fruit flies, worms, zebrafish, and mice.
Why Are Model Organisms Useful? [1]
- Similarity in humans between nervous, cardiovascular, and other biological systems
- Disease time length progressed in animal models
- Cost of maintaining colonies
- Genetic Homology
- Similar phenotypes in humans
- Important fo testing potential therapeutic targets
Examples of Model Organisms for FUS [2]
|
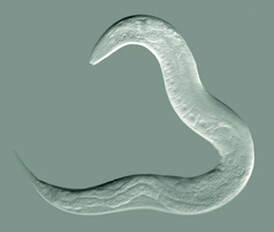
Figure 1: C. elegan
|
Nematode (Caenorhabditis elegans)The C. elegan possess a transparent body capable of growing from embryo to mature adult in only three and half days. In addition, these organisms are hermaphrodites, which makes it easy to create recessive mutations. Most importantly, this model organism has simplicity and is important for analyzing developmental genetic questions due to its exact 959 somatic cells. The cell fat map is the same across the species allowing mutations to be easily identified.
|
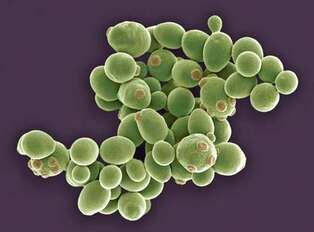
Yeast (Saccharomyces cerevisiae) [3]S. cerevisiae can divide rapidly and in large amount allowing it to be useful for studying metabolic pathways. Although no homolog is present in this species, researchers have utilized GFP-FUS fusions in order to study the disease in this research model. These yeast may not have direct cell to cell communication, but rather do contain conserved machinery, like protein quality-control systems. Thus, making it another model for studying neurodegenerative diseases.
|
Figure 2: S. cerevisiae
|
|
Figure 3: M. musculus
|
Mouse (Mus musculus)Mice have been highly studied with FUS mutations, including other genes that they contain. Transgenic mice are available that contain knockout mutations. These organism are important for analyzing immune and chemical treatments. However, these organisms develop in utero and is hard to analyze impacts on development. In addition, generation time spans around 3 months making screening possible but difficult.
|
Zebrafish (Danio Rerio)Zebrafish have easily observable eggs and embryos for genetic studies. These embryos are transparent and develop outside of the mother's body so any phenotype is easily observable. They also possess a relative small simple genome and are easy to breed in large numbers. Generation time is two to four months, yet have a relatively quick development in early stages. One day after fertilization, many of the vital organs and tissues have been formed. In addition, there are techniques that can be used to easily create recessive mutations that analyze both paternal and maternal effects.
|
Figure 4: D. rerio
|
|
Figure 5: D. melanogaster
|
Fruit Fly (Drosophila melanogaster)The fruit fly has been extensively studied throughout genetic research for behavioral defects, circadian rhythm defects, and understanding early developmental pathways . The fruit fly can easily be grown in the laboratory and has a generation time of only two weeks. Since this organisms has been extensively studied throughout the history of genetic research, vast amounts of information is present on its 13,600 genes and multiple pathways.
|
Model Organism Databases
Model Organism Databases are collections of data on gene ontology, phenotypes, diseases, and expressions of genes in those organisms. This data can be important for determining which phenotype is similar to humans and which aspect of the gene associates with the disease. Some examples of the databases are listed below including:
Flybase [4]
Flybase provided FUS phenotypes observed in Drosophilia showed locomotive behavior defects, increased cell death, and short lived lifespan. Many of these specific phenotypes were observed in muscles and motor neurons of these flies. UAS tissue specific strains with a P398L point mutation showed eye defects in color pigmentation, suggesting cellular death and degeneration.
MGI: Mouse Database [5]
The MGI database provided multiple phenotypes associated with FUS alterations showed increased mortality, behavioral defects, growth size, and neurology defects. One homozygous strain listed showed chromosome breakage, decreased B cell number, and decreased lymphocyte cell number. Another homozygous strain showed abnormal meiosis and increased mortality when exposed to gamma irradiation. As expected, some strains showed motor neuron death and premature death of the organism. The cellular component of FUS was shown to be the nucleus in the mouse and the molecular function of protein binding was similar as in humans. Expression results showed expression of the FUS protein in multiple tissues, including the nervous system.
ZFIN: Zebrafish Database [6]
The Zebrafish Database provided information on multiple strains with specific mutations in the FUS gene. Once again, the molecular function was classified as DNA, RNA, and metal ion binding, due to the presence of the two conserved binding domains. In the zebrafish model, the biological function role was development in axon extension and regulation of transcription. The database provided multiple phenotypes shown in the other species above arising from point, splice, and premature stop mutations.
How Do FUS Mutations Specifically Affect Danio rerio? [6]
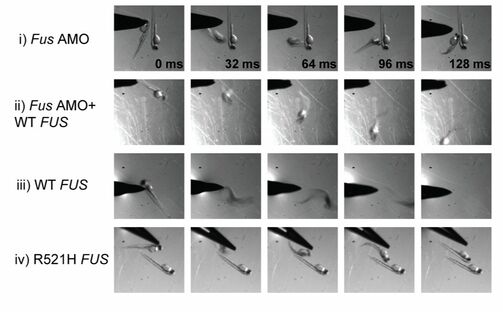
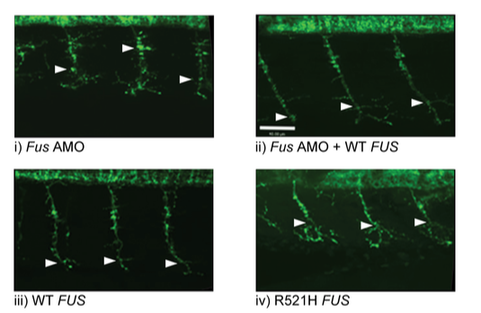
According to ZFIN, there are several phenotypes associated with FUS mutations in ALS. These mutants show decreased lengths of CaP motor neuron axons, axon extension processing quality, decreased FUS expression, impaired thigmotaxis (touch response), fertile organisms, and abnormal larval locomotory behavior. Below are some examples of the phenotypes:
Thigmotaxis Impairement [7]Figure 6: Movement Impairment associated with FUS Mutations
|
Decreased Motor Neuron Length [7]Figure 7: Decreased Motor Length with FUS Mutations
| ||||||
Conclusion
Danio rerio would be a useful model for studying alterations to the FUS gene and analyzing the association with both ALS and DNA repair defects. These organisms are relative cheap, easy to maintain, possess transparent embryos for analyzing aggregate formation, and undergo early stages of development relatively quickly. Full genome sequences are available and this organism has been used previously in other ALS studies. The phenotypes of thigmotaxis impairment and decreased motor length could be an easy way of screening for FUS variants. Their transparent embryo could be useful in order to analyze gene expressions and visualize DNA damage. By using this model organism, it should be relatively efficient to screen for DNA repair defects associated with FUS and ALS. Through these studies, it may be possible to suggest a complete mechanism of DNA repair and discover the underlying cause of ALS.
References
1. Using Research Organisms to Study Health and Disease. (2018, November). Retrieved from https://www.nigms.nih.gov/education/Pages/modelorg_factsheet.aspx
2. Murphy, P. (2013). Lecture 2: Model Organisms. Retrieved from http://virtuallaboratory.colorado.edu/DEVO@CU/papers/pmby1101-lecture2-2013.pdf
3. Kryndushkin, D., & Shewmaker, F. (2011). Modeling ALS and FTLD proteinopathies in yeast: an efficient approach for studying protein aggregation and toxicity. Prion, 5(4), 250-7.
4. Retrieved 3/14/2019 from http://flybase.org/reports/FBgn0285954
5. Retrieved 3/14/2019 from http://www.informatics.jax.org/marker/MGI:1353633
6. Retrieved 3/14/2019 from http://zfin.org/ZDB-GENE-040426-1010
7. Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.
Header Image: https://www.genome.gov/imagegallery/viewimage/?imageID=26454931213
Figure 1: https://images.theconversation.com/files/23633/original/kybzfxcc-1368418075.jpg?ixlib=rb-1.1.0&q=45&auto=format&w=1000&fit=clip
Figure 2: https://www.britannica.com/science/yeast-fungus
Figure 3: https://iiif.elifesciences.org/lax:05959%2Felife-05959-fig2-v1.tif/full/full/0/default.jpg
Figure 4: http://dukemagazine.duke.edu/sites/default/files/pri_img/An%20Interview%20With%20The%20Zebrafish/zebrafish-1200x700.jpg
Figure 5: http://www.animalresearch.info/en/designing-research/research-animals/drosophila-melanogaster/
Figure 6: Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.
Figure 7: Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.
2. Murphy, P. (2013). Lecture 2: Model Organisms. Retrieved from http://virtuallaboratory.colorado.edu/DEVO@CU/papers/pmby1101-lecture2-2013.pdf
3. Kryndushkin, D., & Shewmaker, F. (2011). Modeling ALS and FTLD proteinopathies in yeast: an efficient approach for studying protein aggregation and toxicity. Prion, 5(4), 250-7.
4. Retrieved 3/14/2019 from http://flybase.org/reports/FBgn0285954
5. Retrieved 3/14/2019 from http://www.informatics.jax.org/marker/MGI:1353633
6. Retrieved 3/14/2019 from http://zfin.org/ZDB-GENE-040426-1010
7. Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.
Header Image: https://www.genome.gov/imagegallery/viewimage/?imageID=26454931213
Figure 1: https://images.theconversation.com/files/23633/original/kybzfxcc-1368418075.jpg?ixlib=rb-1.1.0&q=45&auto=format&w=1000&fit=clip
Figure 2: https://www.britannica.com/science/yeast-fungus
Figure 3: https://iiif.elifesciences.org/lax:05959%2Felife-05959-fig2-v1.tif/full/full/0/default.jpg
Figure 4: http://dukemagazine.duke.edu/sites/default/files/pri_img/An%20Interview%20With%20The%20Zebrafish/zebrafish-1200x700.jpg
Figure 5: http://www.animalresearch.info/en/designing-research/research-animals/drosophila-melanogaster/
Figure 6: Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.
Figure 7: Kabashi, E., Bercier, V., Lissouba, A., Liao, M., Brustein, E., Rouleau, G. A., & Drapeau, P. (2011). FUS and TARDBP but not SOD1 interact in genetic models of amyotrophic lateral sclerosis. PLOS Genetics, 7(8). doi: 10.1371/journal.pgen.1002214.