What is Gene Ontology? [1]
The Gene Ontology Consortium describes gene ontology as the network of biological classes that represent all known aspects of biology in the natural world. These networks help establish genes by compartmentalizing them into three different categories: by their Cellular Compartment, Molecular Functions, and Biological Processes.
Cellular Compartment: This is described as the component of cell or area in a cell were the gene performs a function. This can be an anatomical structure (like the nucleus) or macromolecular complexes (like the ribosome). It is important to note that this category refers to the cellular anatomy rather than the specific process.
Molecular Function: These are activities performed by products of a gene on a smaller level. These describe roles at the molecular level like transport. The terms represent these roles than the specific molecules that perform the tasks. These terms do not give specifics where the process takes place. These functions are commonly referred to as an "activity" and can be broad or specific. One molecular function example would be transporter activity.
Biological Processes: This is referred to as the larger process accomplished by multiple molecular functions. These can be general broad biological processes like DNA repair or more specific molecular activities. Again, it is important to remember that these are molecular functions and not pathways or dynamics.
Cellular Compartment: This is described as the component of cell or area in a cell were the gene performs a function. This can be an anatomical structure (like the nucleus) or macromolecular complexes (like the ribosome). It is important to note that this category refers to the cellular anatomy rather than the specific process.
Molecular Function: These are activities performed by products of a gene on a smaller level. These describe roles at the molecular level like transport. The terms represent these roles than the specific molecules that perform the tasks. These terms do not give specifics where the process takes place. These functions are commonly referred to as an "activity" and can be broad or specific. One molecular function example would be transporter activity.
Biological Processes: This is referred to as the larger process accomplished by multiple molecular functions. These can be general broad biological processes like DNA repair or more specific molecular activities. Again, it is important to remember that these are molecular functions and not pathways or dynamics.
GO Terms of FUS
Cellular Component [2]
|
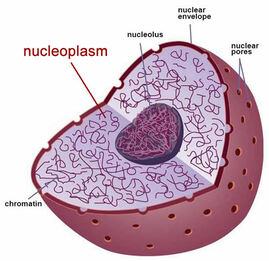
Wild-type FUS is largely localized in the nucleus, but more specifically the nucleoplasm. These locations are shown in on the right (Figure 1). Other articles have suggested FUS to be active in the polysome (cluster of ribosomes), perikaryon (cell body of neuron containing the nucleus), dendritic spine head, and perinuclear region of the cytoplasm. In contrast to these normal locations, mutant FUS tends to localize in the cytoplasm and form aggregates that are no longer able to travel back to the nucleus. This may suggest that FUS is no longer able to perform normal functions in the nucleus. This disruption may suggest multiple mechanisms that may be altered.
|
Figure 1: Schematic Drawing of the Nucleus
|
Molecular Function [2]
|
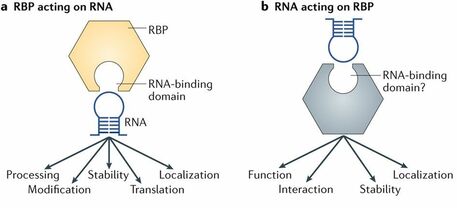
Figure 2: Depiction of the RNA-Binding Domain
|
FUS plays a normal role in DNA, RNA, protein and chromatin binding. FUS has two protein domains that are a zinc-finger domain and RNA binding motif. These two domains are able to participate in these binding relationships. In addition, FUS has been suggested to play a role binding to ionictropic glutamate, thyroid hormone, estrogen receptors, and retanoid x receptors. Figure 2 shows the molecular function of RNA binding at the top of the figure, while the bottom arrows suggest biological functions due to these molecular functions.
|
Biological Processes [2]
|
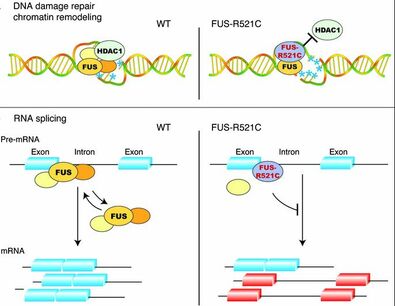
Due to the role of FUS in RNA binding, it is also involved with the regulation of both transcriptional and splicing. This primarily occurs through the interactions of the RNA polymerase II and the spliceosome respectively. This protein has also been involved with the promotion of DNA repair mechanisms and facilitating translation. It has been known that FUS is associated with repairing both single strand breaks (SSBs) and double strand breaks (DSBs) via known DNA repair mechanisms. Primarily it has been shown to interact with PARP1, LigIII, and XRCC1 in SSBs. The downstream repair mechanisms for SSBs/DSBs and are relatively unknown in FUS mutants. By understanding which interactions are altered, it could lead to reasons why we see early cases of death in FUS-ALS individuals.
|
Figure 3: Biological Processes Impacted by FUS Mutations
|
Conclusion
By understanding the Gene Ontology of FUS, you are better able to understand the role of FUS in ALS, neurodegeneration, and DNA repair. The molecular location of FUS in those with non variant alleles provides an idea of what other roles could be carried about by the products of these genes. In addition, the molecular location and biological processes associated with the FUS protein can highlight what specific function is being altered in these cases. From these findings, we can better try to understand the specific cause of ALS in these cases and discover what aspect of this multi-functional protein is leading to the death of these motor neurons.
References
[1]. Gene Ontology overview. (n.d.). Retrieved from http://geneontology.org/docs/ontology-documentation/
[2]. RNA Binding FUS. (n.d.) Retrieved from http://amigo.geneontology.org/amigo/gene_product/UniProtKB:P35637
Header Image: https://brooklynrail.org/article_image/image/20603/Taney_3.jpg
Figure 1: https://2.bp.blogspot.com/-bSF5Z-Bz8wQ/WaVjy1jiXvI/AAAAAAAAj_E/9bVh6UBbX8IYEkAwM1D3uebslu6ZZJQgACLcBGAs/s1600/nucleus.jpg
Figure 2: https://media.springernature.com/full/nature-static/assets/v1/image-assets/nrm.2017.130-f1.jpg
Figure 3: https://www.researchgate.net/profile/Eric_Huang10/publication/299475594/figure/fig4/
AS:392331896344576@1470550621059/Mechanisms-of-wild-type-and-mutant-FUS-in-DNA-damage-repair-response-and-RNA-splicing.png
[2]. RNA Binding FUS. (n.d.) Retrieved from http://amigo.geneontology.org/amigo/gene_product/UniProtKB:P35637
Header Image: https://brooklynrail.org/article_image/image/20603/Taney_3.jpg
Figure 1: https://2.bp.blogspot.com/-bSF5Z-Bz8wQ/WaVjy1jiXvI/AAAAAAAAj_E/9bVh6UBbX8IYEkAwM1D3uebslu6ZZJQgACLcBGAs/s1600/nucleus.jpg
Figure 2: https://media.springernature.com/full/nature-static/assets/v1/image-assets/nrm.2017.130-f1.jpg
Figure 3: https://www.researchgate.net/profile/Eric_Huang10/publication/299475594/figure/fig4/
AS:392331896344576@1470550621059/Mechanisms-of-wild-type-and-mutant-FUS-in-DNA-damage-repair-response-and-RNA-splicing.png
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.